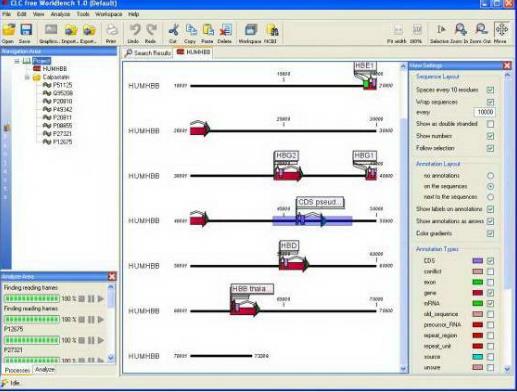
The first primer pair seems very well, we are going to settle for this pair. Thus a large deviation from the ideal value and a small tolerance will give a large negative contribution in the final score and a small deviation from the ideal and a high tolerance will give a small negative contribution in the final score. The negative contribution to the final score is determined by how much the parameter deviates from the ideal value and is scaled by the specified tolerance:Ĭontribution = (ideal - actual value)/tolerance Self-annealing score: the ideal value is 0 and the tolerance corresponds to the maximum value specified in Each parameter is assigned an ideal value and a tolerance. The ideal score for a pair is 100 and pairs are ranked inĭescending order. Primer pairs such as the pair-annealing score. Parameters related to individual primers, such as the secondary structure score, and parameters related to FAST protein experiment templates (from Flies et al.CLC Main Workbench employs a proprietary algorithm to rank primer pairs.FAST protein protocol (from Flies et al., Science Advances, in press).Protein alignments and general protein and DNA analysis: CLC Sequence Viewer (free version)įree software environment for statistical computing and graphics: R-project, RStudioįree image editing software: GNU Image Manipulation Program (GIMP)įree software to design degenerate primers: HYDENīuffer preparation calculator from CHO cell adaption to serum-free media 3.2.2 Analysis of the NGS data using CLC Genomics Workbench software. Predict nuclear localization signals: cNLS mapperįind potential restriction sites in a DNA sequence by introducing silent mutations that do not change the protein sequence: Webcutter or SilentMutatorĪmino acid properties and consequences of substitutions: Russel LabĮxcellent software for plasmid construction, sequence analysis and general purpose DNA work: SnapGene Predict non-classically secreted proteins: SecretomePĬalculate multi-protein percent identity: Clustal Omega Protease Specificity Prediction Server: iProt-Sub Predict molecular weight and other good stuff: Protparam ProtPI Predict protein domains and signal peptides: Phobius 
Predict protein structure and features: SMART (e.g. Predict Eukaryotic Linear Motifs (ELM) (e.g. Plasmids 101: The promotor region (Addgene) Download PDFīetter Plasmid Midipreps Part II: What Causes Low Yields? (BitesizeBio) Making LB agar plate – protocol Making LB agar plate – video (Addgene)Īntibiotic concentrations spreadsheet (Addgene, InvivoGen)Īntibiotic concentrations (Thermo Fisher) Plasmid guide (Addgene) – very useful for cloning questions Interactive molecular biology tools (NEB) Ten Tips for pipetting a 384-well plate (BitesizeBio) Useful Numbers for Cell Culture (Thermo Scientific) Landscape of Immuno-Oncology Drug Development interactive tool for keeping up with the development of anti-cancer drugs.Ĭell Counting with a Hemocytometer (BitesizeBio) – Click here to download PDF The Biolgend web is an outstanding for all things immunologyįluorescence spectra analyzer (Biolegend) Immuno-Navigator A database for gene coexpression in the immune system Generation and Testing of Fluorescent Adaptable Simple Theranostic (FAST) Proteins (peer-reviewed Bio-protocol methods used in Flies et al., 2020 Science Advances)įAST protein recipes (from Flies et al., 2019 bioRxiv)įixation and flow cytometry from BitesizeBioīuffer preparation calculator from a great source for find tissue-specific gene expression. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
